O’NEAL
New 1,000-year earthquake simulations suggest parts of the San Andreas system are under record levels of strain
For over a century, Southern California has avoided the catastrophic earthquakes geologists long considered inevitable. However, a groundbreaking computational study suggests that the region is now under tectonic stress levels unseen in the past millennium. This discovery was made possible by advanced earthquake-cycle simulations that reconstruct 1,000 years of fault behavior.
A collaborative team from the University of Bern, the University of Hawaiʻi at Mānoa, Northern Arizona University, the U.S. Geological Survey, and the Scripps Institution of Oceanography developed a sophisticated four-dimensional model of the Southern San Andreas Fault System. Their findings reveal that stress has reached critical, historically high levels near Cajon Pass, a pivotal junction where the San Andreas and San Jacinto faults intersect.
Published in the Journal of Geophysical Research: Solid Earth, this research arrives amid rising concerns that Southern California may be nearing a significant seismic event. As the study notes, the region is essentially sitting on a fault system that has been storing energy for generations; following the massive 7.9 magnitude Fort Tejon earthquake of 1857, the southern San Andreas has remained uncharacteristically quiet.
A thousand years of earthquakes reconstructed in silicon
The study’s most significant contribution is not simply its conclusions, but the computational machinery used to reach them.
Researchers employed a physics-based earthquake-cycle simulator known as Maxwell, a semi-analytic Fourier-transform model capable of tracking stress evolution across hundreds of kilometers of fault networks. The model represents an elastic crust resting atop a viscoelastic mantle and calculates how tectonic loading, earthquake ruptures, and post-seismic relaxation interact through time.
The computational domain spans approximately 450 by 900 kilometers and incorporates 38 major fault segments extending from California’s Carrizo Plain to the Borrego Mountain region. The simulation integrates geodetic observations, fault locking depths, geological slip rates, and a detailed paleoseismic record covering the last millennium.
Rather than examining a single earthquake, the researchers recreated roughly 1,000 years of earthquake history, modeling dozens of large ruptures and tracking how stress accumulated and transferred between interconnected fault segments over centuries. The resulting calculations generated time-dependent stress fields at 10-year intervals, with annual-resolution simulations around major earthquake events.
This type of multi-century earthquake-cycle modeling would have been impossible only a few decades ago. The calculations involve repeated evaluations of three-dimensional fault dislocations, viscoelastic relaxation processes, and Coulomb stress interactions across an entire regional fault network.
Cajon Pass: California’s potential earthquake gate
At the center of the investigation lies Cajon Pass, a narrow corridor northeast of Los Angeles where two of California’s most important fault systems converge.
The location carries extraordinary significance because it appears capable of acting as what researchers call an “earthquake gate.” Under some stress conditions, ruptures stop at the junction. Under others, they may pass through it and cascade into neighboring faults, producing substantially larger earthquakes.
Historical evidence hints at both possibilities.
The massive 1857 Fort Tejon earthquake appears to have terminated near Cajon Pass. In contrast, an earlier 1812 earthquake may have propagated through the junction, linking multiple fault segments in a much larger interconnected rupture.
The new simulations suggest that stress relationships between neighboring fault segments may determine whether the gate remains closed or swings open.
Researchers found that when stress levels on the San Andreas and San Jacinto systems become more closely aligned, through-going ruptures become more likely. This raises the possibility that future earthquakes could involve multiple fault systems simultaneously rather than remaining confined to a single segment.
For Southern California’s densely populated urban corridor, that distinction matters enormously.
Stress levels reach modern extremes
The most troubling findings emerge from the model’s estimate of present-day conditions.
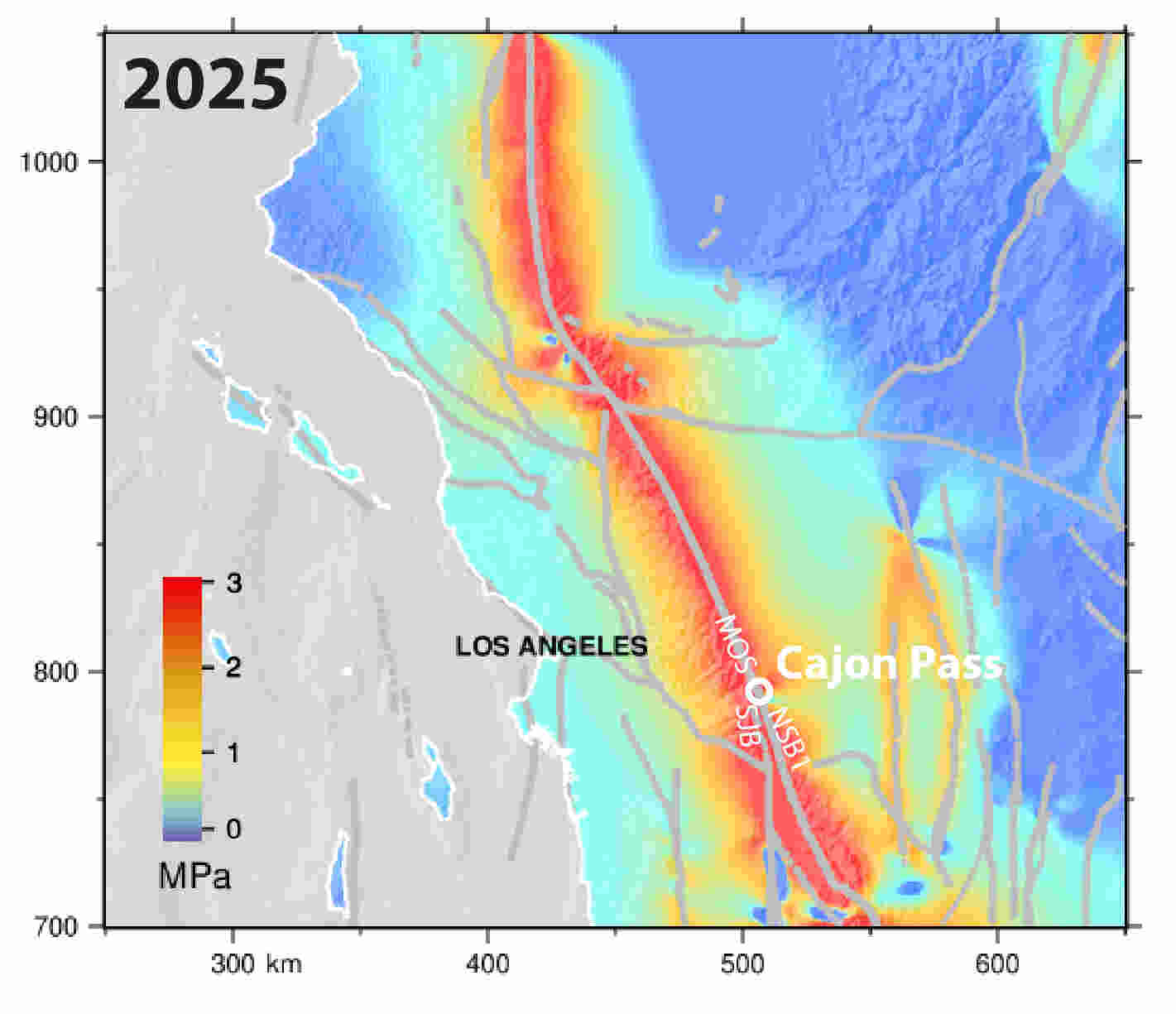
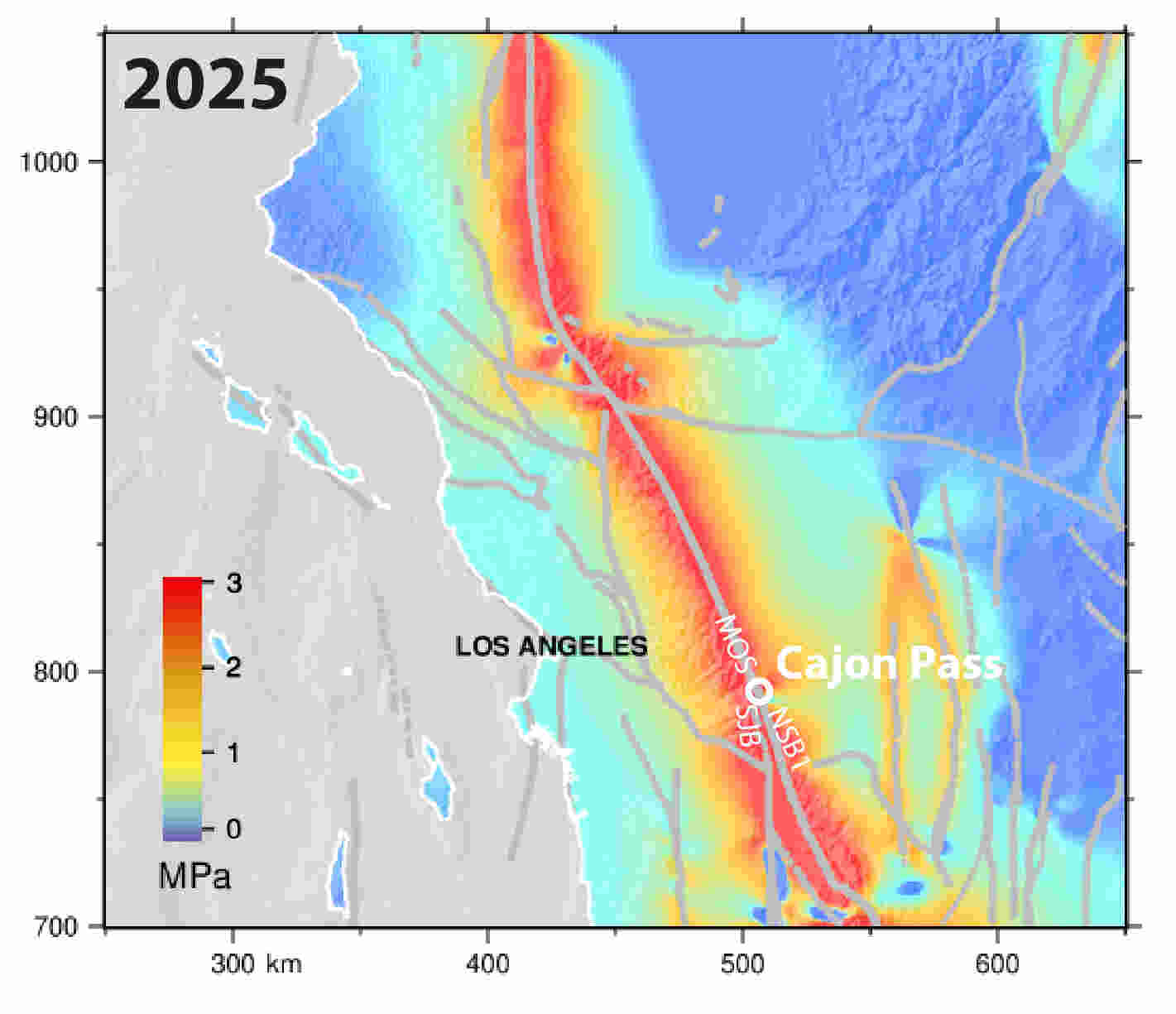
The simulations indicate that tectonic stress has accumulated steadily since the nineteenth century, producing elevated stress levels throughout the region. By the model’s 2025 endpoint, the Mojave South segment of the San Andreas carried approximately 2.8 megapascals of Coulomb stress, while the San Jacinto Bernardino segment reached roughly 3.6 megapascals. The latter exceeds any stress level modeled on that segment during the previous millennium.
The Mojave South segment is particularly concerning because researchers identified it as carrying the highest stress accumulation rate in the system, approximately 1.8 megapascals per century.
Even more striking is the historical context.
The study reports that current stress on the Mojave South segment is the highest observed anywhere in its modeled 1,000-year history. Meanwhile, stress levels on the San Jacinto Bernardino segment now exceed those present before any major rupture included in the simulation record.
Although the researchers emphasize that these values should not be interpreted as direct earthquake predictions, they nevertheless portray a fault system that has been loading for an exceptionally long period.
Why supercomputing matters
Earthquake hazards are notoriously difficult to forecast because faults do not operate independently.
A rupture on one fault can increase stress on neighboring segments while reducing stress elsewhere. These interactions can persist for decades or centuries. Untangling those relationships requires simulations that capture the evolution of entire fault networks rather than isolated faults.
The present study demonstrates how high-performance computing is transforming seismic hazard science from a largely observational discipline into a predictive modeling enterprise.
By assimilating paleoseismic records, geological slip rates, geodetic measurements, and rheological models into a unified computational framework, researchers can now evaluate thousands of years of fault evolution and explore scenarios that have never been directly observed.
The team even simulated hypothetical future rupture sequences, including scenarios where multiple fault segments fail together. These experiments revealed that a complete rupture involving all major Cajon Pass fault strands would produce the largest stress release observed in the entire modeled history.
Such analyses are increasingly relevant as emergency planners, infrastructure operators, and policymakers seek more sophisticated estimates of seismic risk.
The uncomfortable message
The study stops well short of forecasting an imminent earthquake. Earthquake occurrence remains fundamentally unpredictable, and the authors repeatedly caution that their results depend on model assumptions and fault parameters.
Yet the broader message is difficult to ignore.
More than 169 years have passed since the Fort Tejon earthquake ruptured the southern San Andreas. During that time, plate tectonic motion has continued relentlessly, loading the fault system year after year.
The simulations indicate that stress levels today rival, or in some cases exceed, those that preceded major historical ruptures.
For residents of Southern California, that is a sobering conclusion.
For the supercomputing community, however, it is also evidence of a profound shift in Earth science.
The most important discoveries about future earthquake hazards may no longer come solely from the ground beneath our feet, but from the massive computational systems capable of reconstructing centuries of tectonic history and revealing what the Earth’s faults have been quietly storing all along.








 How to resolve AdBlock issue?
How to resolve AdBlock issue?