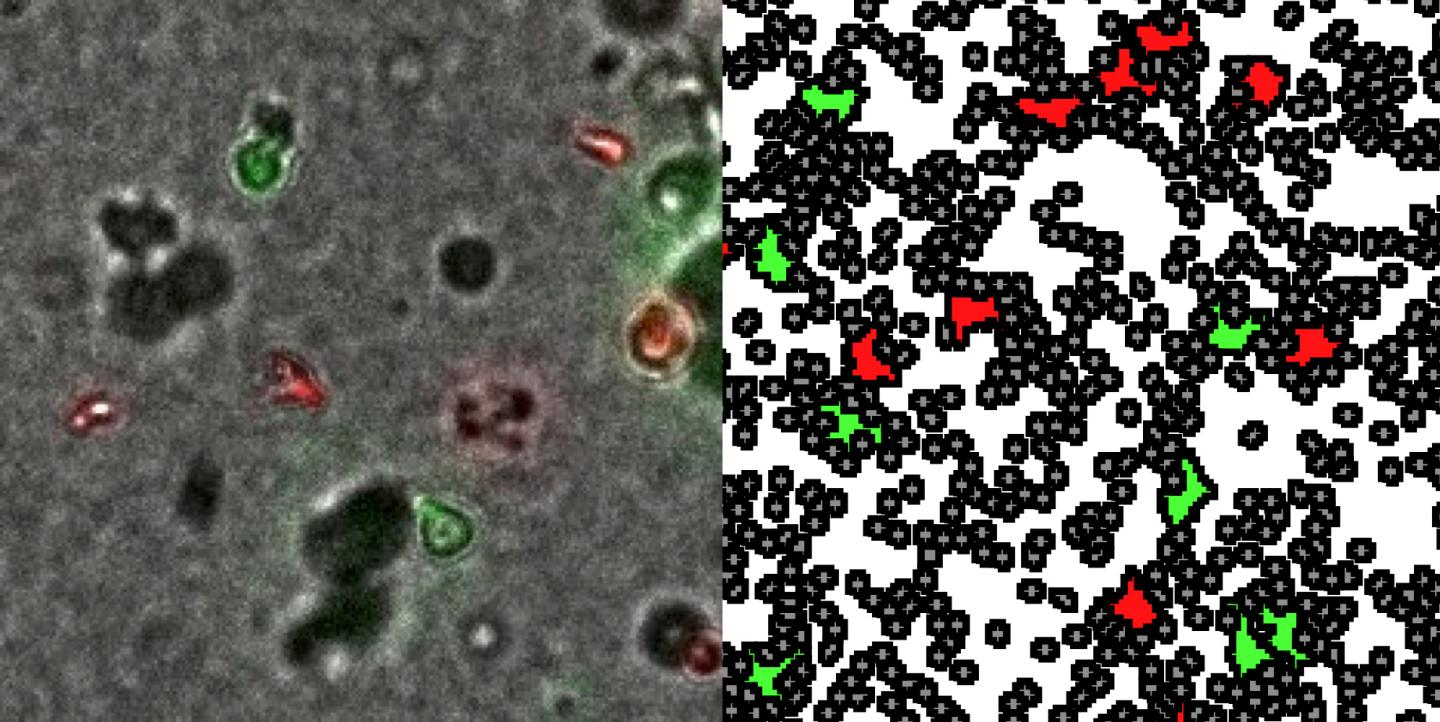
Researchers at Dartmouth's Norris Cotton Cancer Center have created a deep learning model that can accurately identify cancerous esophagus tissue on microscopy images without the time-consuming manual data input required for current methods
Recently, deep learning methods have shown promising results for analyzing histological patterns in microscopy images. These approaches, however, require a laborious, high-cost, manual annotation process by pathologists called "region-of-interest annotations." A research team at Dartmouth and Dartmouth-Hitchcock Norris Cotton Cancer Center, led by Saeed Hassanpour, Ph.D., has addressed this shortcoming of current methods by developing a novel attention-based deep learning method that automatically learns clinically important regions on whole-slide images to classify them.
The team tested their new approach for identifying cancerous and precancerous esophagus tissue on high-resolution microscopy images without training on region-of-interest annotations. "Our new approach outperformed the current state-of-the-art approach that requires these detailed annotations for its training," concludes Hassanpour. Their results, "Detection of Cancerous and Precancerous Esophagus Tissue on Histopathology Slides Using Attention-Based Deep Neural Networks" will publish in JAMA Network Open in early November 2019. {module INSIDE STORY}
For histopathology image analysis, the manual annotation process typically outlines the regions of interest on a high-resolution whole slide image to facilitate training the computer model. "Data annotation is the most time-consuming and laborious bottleneck in developing modern deep learning methods," notes Hassanpour. "Our study shows that deep learning models for histopathology slides analysis can be trained with labels only at the tissue level, thus removing the need for high-cost data annotation and creating new opportunities for expanding the application of deep learning in digital pathology."
The team proposed the network for Barrett esophagus and esophageal adenocarcinoma detection and found that its performance exceeds that of the existing state-of-the-art method. "The result is significant because our method is based solely on tissue-level annotations, unlike existing methods that are based on manually annotated regions," says Hassanpour. He expects this model to open new avenues for applying deep learning to digital pathology. "Our method would facilitate a more extensive range of research on analyzing histopathology images that were previously not possible due to the lack of detailed annotations. Clinical deployment of such systems could assist pathologists in reading histopathology slides more accurately and efficiently, which is a critical task for the cancer diagnosis, predicting prognosis, and treatment of cancer patients."
Looking ahead, Hassanpour's team is planning to validate their model further by testing it on data from other institutions and running prospective clinical trials. They also plan to apply the proposed model to histological images of other types of tumors and lesions for which training data are scarce or bounding box annotations are not available.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module In article}
{module In article}