Magnetic vortices could be manipulated by waste heat to realize low-power supercomputing applications
Tiny amounts of heat can be used to control the movement of magnetic whirlpools called skyrmions, RIKEN physicists have shown. This ability could help to develop energy-efficient forms of supercomputing that harness waste heat. 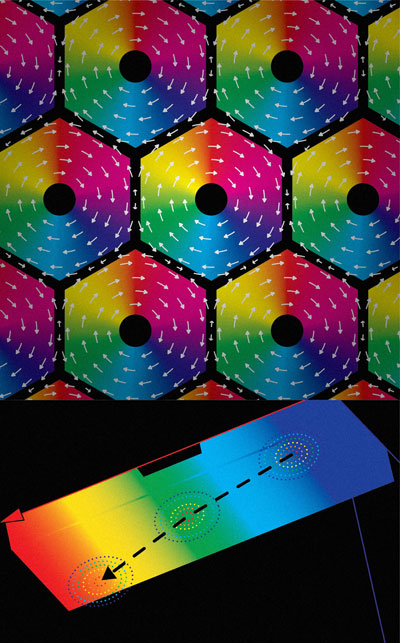
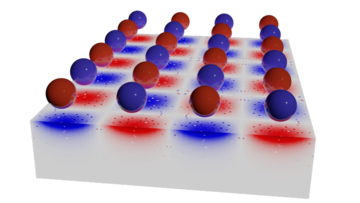
Skyrmions are minuscule vortices that form when the magnetic flux of a group of atoms organizes into swirling patterns. Skyrmions can move around inside a material, and under certain conditions, they cluster together to form a regular arrangement known as a skyrmion lattice (upper part of Fig. 1).
Skyrmions are promising information carriers in next-generation computer chips that have very low power requirements. Researchers can already control skyrmions by applying electrical currents and magnetic fields, but they are seeking to manipulate them using heat flow instead. “This is an exciting prospect since it would raise the possibility of using waste heat to move skyrmions around,” says Xiuzhen Yu at the RIKEN Center for Emergent Matter Science.
Now, Yu and her colleagues have shown how a temperature gradient can be used to propel skyrmions in an electrically insulating magnetic material.
The team built a device that consisted of a plate of this material, a miniature heating element, and two electric thermometers. They then generated skyrmions that were roughly 60 nanometers wide in the plate by cooling it to about −253 degrees Celsius and applying a magnetic field. These skyrmions gathered into a stable honeycomb structure known as a hexagonal skyrmion lattice.
Yu’s team then increased the temperature slightly at one end of the plate and used a transmission electron microscope to watch how this affected the skyrmions. A temperature gradient of 100th of a degree per millimeter of the plate was enough to nudge the skyrmions into motion. Above this threshold, the edge of the honeycomb lattice drifted from the cooler to the warmer end of the plate, traveling in the opposite direction to the flow of heat (lower part of Fig. 1). This required a very low heat power of just 10 microwatts, which is hundreds or thousands of times smaller than the power needed to move skyrmions using electrical currents or magnetic fields. Using a slightly higher power, individual skyrmions could be driven through the plate by the temperature gradient.
The researchers say that this is the first time that heat-driven skyrmion motion has been seen in an insulating magnet. “This finding should stimulate researchers to develop energy-efficient devices by using skyrmions,” says Yu.
The team is now studying the heat-induced dynamics of skyrmions, including their transformation into their anti-particles—anti-skyrmions in metallic systems at room temperature.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 


