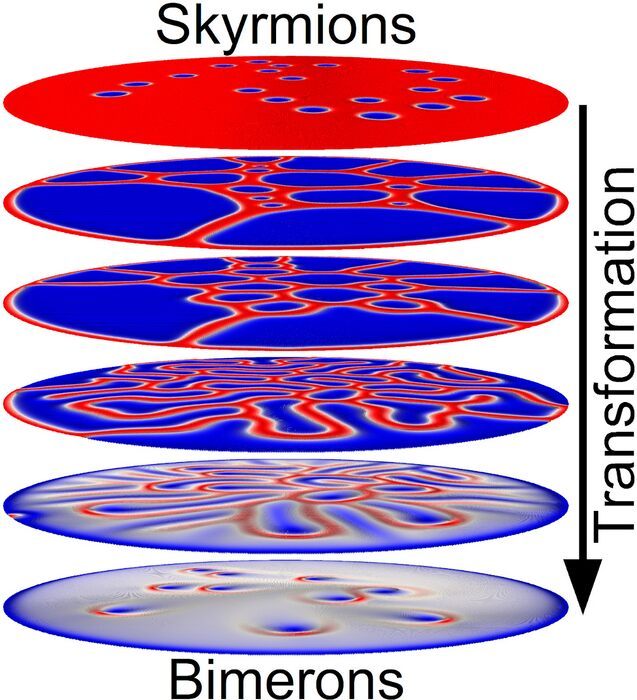
 The transformation between skyrmions and bimerons has now been realized by scientists at Japan's Shinshu University
The transformation between skyrmions and bimerons has now been realized by scientists at Japan's Shinshu University
Skyrmions and bimerons are fundamental topological spin textures in magnetic thin films with asymmetric exchange interactions and they can be used as information carriers for next generation low energy consumption memory, advanced neuromorphic supercomputing, and advanced quantum supercomputing as they have multiple degrees of freedom that can carry information. The transformation between isolated skyrmions and bimerons will be an essential operation for future computing architecture based on multiple different topological bits. Therefore, the community needs to find effective ways to realize the creation, transformation, and manipulation of skyrmions and bimerons in magnetic materials.
In a recent study published in Nano Letters, the group led by Xiaoxi Liu, a Professor in the Department of Electrical and Computer Engineering at Shinshu University in Japan and their international collaborators demonstrate in experiments and simulations that the creation of isolated skyrmions and their subsequent transformation to bimerons are possible in a magnetic disk surrounded by a current-carrying and omega-shaped microcoil, where the electric current-induced Oersted field and temperature-induced perpendicular magnetic anisotropy variation play important roles in the transformation between skyrmions and bimerons. Researchers find that the current injected into the microcoil can generate an Oersted field to switch the magnetization of the magnetic disk in the out-of-plane directions. Meanwhile, the current injected into the microcoil can heat the magnetic disk and increases the device's temperature. As a result, a temperature-induced decrease of magnetic anisotropy is realized in the magnetic disk, which leads to the magnetization reorientation from the out-of-plane direction to the in-plane direction and thus, fosters the transformation from skyrmions to bimerons. Researchers also find deformed skyrmion bubbles and chiral labyrinth domains during the transformation between skyrmions and bimerons.
The researchers’ results demonstrate the possibility that two different types of topological spin textures can be hosted by the same magnetic film with asymmetric exchange interactions, which may provide guidelines for building novel spintronic applications based on different types of topological spin textures.
“Our experiment clarified for the first time the transformation between different topological spin textures,” explains Liu. He also mentions, “Skyrmions and bimerons are two most important information carriers for next-generation memory and advanced computing architectures. Our research has a fundamental physical interest. It is also important for future data storage and computing community”.
Researchers will try to study magnetic and spintronic device applications based on the transformation of different types of topological spin textures. An example is voltage-gated spintronic devices based on skyrmions and bimerons. “Our ultimate goal is the application of topological spin textures for low energy consumption, high-density memory, and advanced neuromorphic computing,” says Liu.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?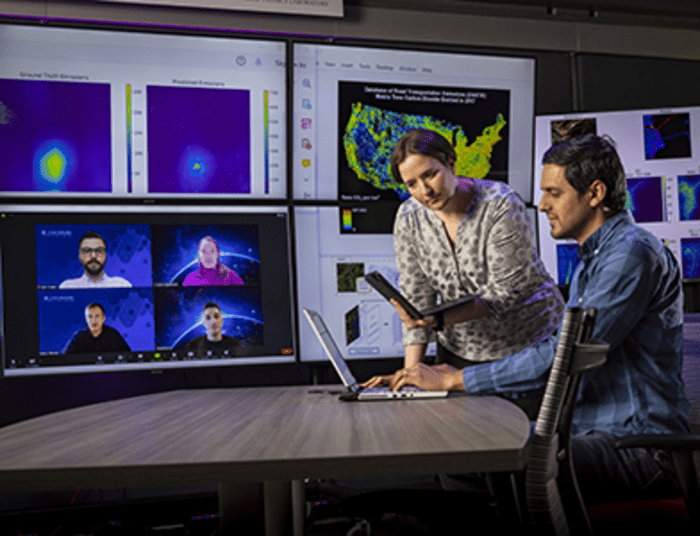  The Environmental Protection Agency estimates that the transportation sector accounts for approximately 27% of all greenhouse gas emissions annually in the United States, and emissions from road transportation — driven by carbon-creating internal combustion vehicles — account for a large majority of that.
The Environmental Protection Agency estimates that the transportation sector accounts for approximately 27% of all greenhouse gas emissions annually in the United States, and emissions from road transportation — driven by carbon-creating internal combustion vehicles — account for a large majority of that. A supernova is a stellar explosion, which occurs when the lives of some really massive stars come to an end. In this violent epilogue, the star expels the material from its outer layers by means of a shock wave, allowing us to see the various elements it was composed of.
A supernova is a stellar explosion, which occurs when the lives of some really massive stars come to an end. In this violent epilogue, the star expels the material from its outer layers by means of a shock wave, allowing us to see the various elements it was composed of. We breathe to survive. But a breath of fresh air does more than fill our lungs. New research from Aarhus University the second largest and second oldest university in Denmark, indicates that breathing impacts our emotions, attention and how we can process the outside world.
We breathe to survive. But a breath of fresh air does more than fill our lungs. New research from Aarhus University the second largest and second oldest university in Denmark, indicates that breathing impacts our emotions, attention and how we can process the outside world.