Scientists use sophisticated supercomputer simulations and observations of trans-Neptunian objects to understand the formation of the solar system
Using sophisticated supercomputer simulations and observations, a team led by researchers from the Earth-Life Science Institute (ELSI) at Tokyo Institute of Technology has shown how the so-called trans-Neptunian Objects (or TNOs) may have formed. TNOs, which include the dwarf planet Pluto, are a group of icy and rocky small bodies--smaller than planets but larger than comets--that orbit the Solar System beyond the planet Neptune. TNOs likely formed at the same time as the Solar System, and understanding their origin could provide important clues as to how the entire Solar System originated.  {module In-article}
{module In-article}
Like many solar system bodies, including the Earth, TNOs often have their own satellites, which likely formed early on from collisions among the building blocks of the Solar System. Understanding the origin of TNOs along with their satellites may help understand the origin and early evolution of the entire Solar System. The properties of TNOs and their satellites--for example, their orbital properties, composition and rotation rates--provide a number of clues for understanding their formation. These properties may reflect their formation and collisional history, which in turn may be related to how the orbits of the giant planets Jupiter, Saturn, Neptune, and Uranus changed over time since the Solar System formed.
The New Horizons spacecraft flew by Pluto, the most famous TNO, in 2015. Since then, Pluto and its satellite Charon have attracted a lot of attention from planetary scientists, and many new small satellites around other large TNOs have been found. In fact, all known TNOs larger than 1000 km in diameter are now known to have satellite systems. Interestingly, the range of the estimated mass ratio of these satellites to their host systems ranges from 1/10 to 1/1000, encompassing the Moon-to-Earth mass ratio (~1/80). This may be significant because Earth's Moon and Charon are thought to have formed from a giant impactor. 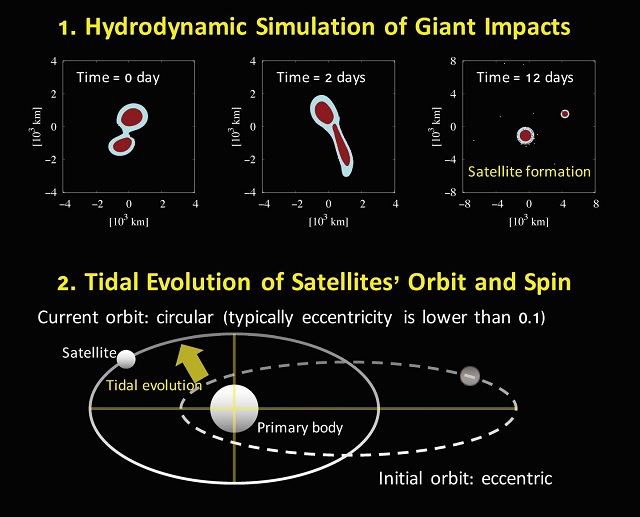 {module In-article}
{module In-article}
To study the formation and evolution of TNO satellite systems, the research team performed more than 400 giant impact simulations and tidal evolution calculations. "This is really hard work," says the study's senior author, Professor Hidenori Genda from the Earth-Life Science Institute (ELSI) at Tokyo Institute of Technology. Other Tokyo Tech team members included Sota Arakawa and Ryuki Hyodo.
The Tokyo Tech study found that the size and orbit of the satellite systems of large TNOs are best explained if they formed from impacts of molten progenitors. They also found that TNOs which are big enough can retain internal heat and remain molten for a span of only a few million years; especially if their internal heat source is short-lived radioactive isotopes such as Aluminum-26, which has also been implicated in the internal heating of the parent bodies of meteorites. Since these progenitors would need to have a high short-lived radionuclide content in order to be molten, these results suggest that TNO-satellite systems formed before the outward migration of the outer planets, including Neptune, or in the first ~ 700 million years of Solar System history.
Previous planet formation theories had suggested the growth of TNOs took much longer than the lifetime of short-lived radionuclides, and thus TNOs must not have been molten when they formed. These scientists found, however, that rapid TNO formation is consistent with recent planet formation studies which suggest TNOs formed via accretion of small solids to pre-existing bodies. The rapid formation of large TNOs is consistent with recent planet formation studies; however, other analyses suggest comets formed well after most short-lived radionuclides had decayed. Thus the authors note that there is still much work to be done to produce a unified model for the origin of the Solar System's planetary bodies. 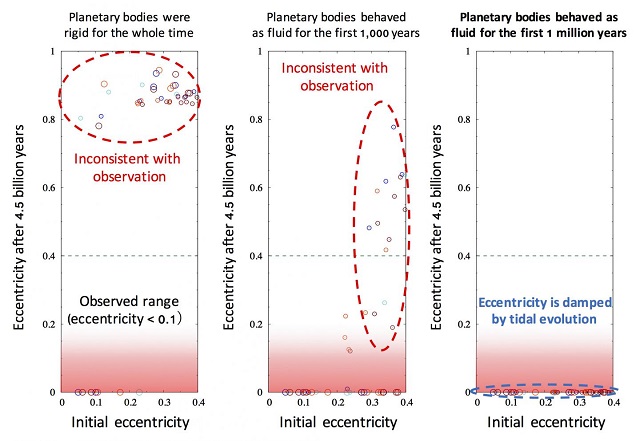 {module In-article}
{module In-article}


 How to resolve AdBlock issue?
How to resolve AdBlock issue?