When pondering the probability of discovering technologically advanced extraterrestrial life, the question that often arises is, "if they're out there, why haven't we found them yet?" And often, the response is that we have only searched a tiny portion of the galaxy. Further, algorithms developed decades ago for the earliest digital computers can be outdated and inefficient when applied to modern petabyte-scale datasets. Now, research led by an undergraduate student at the University of Toronto, Peter Ma, along with researchers from the SETI Institute, Breakthrough Listen, and scientific research institutions around the world, has applied a deep learning technique to a previously studied dataset of nearby stars and uncovered eight previously unidentified signals of interest.
“In total, we had searched through 150 TB of data of 820 nearby stars, on a dataset that had previously been searched through in 2017 by classical techniques but labeled as devoid of interesting signals," said Peter Ma, lead author. “We're scaling this search effort to 1 million stars today with the MeerKAT telescope and beyond. We believe that work like this will help accelerate the rate we’re able to make discoveries in our grand effort to answer the question ‘are we alone in the universe?’”
The search for extraterrestrial intelligence (SETI) looks for evidence of extraterrestrial intelligence originating beyond Earth by trying to detect technosignatures, or evidence of technology, that alien civilizations could have developed. The most common technique is to search for radio signals. Radio is a great way to send information over the incredible distances between the stars; it quickly passes through the dust and gas that permeate space, and it does so at the speed of light (about 20,000 times faster than our best rockets). Many SETI efforts use antennas to eavesdrop on any radio signals aliens might be transmitting.
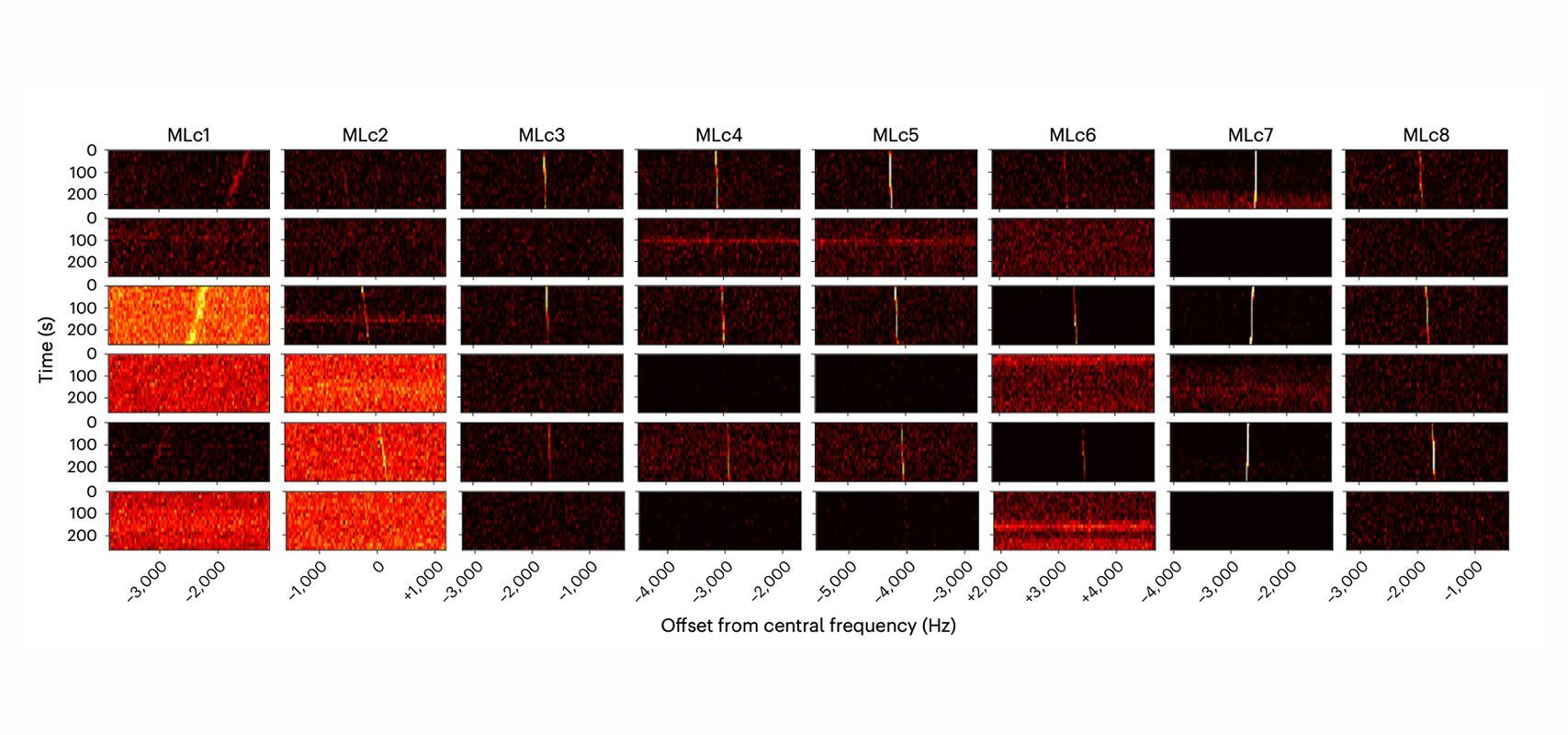
This study re-examined data taken with the Green Bank Telescope in West Virginia as part of a Breakthrough Listen campaign that initially indicated no targets of interest. The goal was to apply new deep learning techniques to a classical search algorithm to yield faster, more accurate results. After running the new algorithm and manually re-examining the data to confirm the results, newly detected signals had several key characteristics:
- The signals were narrow band, meaning they had narrow spectral width, on the order of just a few Hz. Signals caused by natural phenomena tend to be broadband.
- The signals had non-zero drift rates, which means the signals had a slope. Such slopes could indicate a signal’s origin had some relative acceleration with our receivers, hence not local to the radio observatory.
- The signals appeared in ON-source observations and not in OFF-source observations. If a signal originates from a specific celestial source, it appears when we point our telescope toward the target and disappears when we look away. Human radio interference usually occurs in ON and OFF observations due to the source being close by.
Cherry Ng, another of Ma’s research advisors and an astronomer at both the SETI Institute and the French National Center for Scientific Research said, “These results dramatically illustrate the power of applying modern machine learning and computer vision methods to data challenges in astronomy, resulting in both new detections and higher performance. Application of these techniques at scale will be transformational for radio techno signature science.”
While re-examinations of these new targets of interest have yet to result in re-detections of these signals, this new approach to analyzing data can enable researchers to more effectively understand the data they collect and act quickly to re-examine targets. Ma and his advisor Dr. Cherry Ng are looking forward to deploying extensions of this algorithm on the SETI Institute’s COSMIC system.
Since SETI experiments began in 1960 with Frank Drake’s Project Ozma at the Greenbank Observatory, a site now home to the telescope used in this latest work, technological advances have enabled researchers to collect more data than ever. This massive volume of data requires new computational tools to process and analyze that data quickly to identify anomalies that could be evidence of extraterrestrial intelligence. This new machine-learning approach is breaking new ground in the quest to answer the question, “are we alone?”


 How to resolve AdBlock issue?
How to resolve AdBlock issue?